set.seed(1)
no.days <- 14
v.non.smoker <- c(rpois(n = no.days - 1, lambda = 0), 1)
v.smoker1_2 <- rpois(n = no.days, lambda = 5)
v.smoker.box <- rpois(n = no.days, lambda = 20)
d.smokers <- data.frame(no.cigarettes =
c(v.non.smoker, v.smoker1_2, v.smoker.box),
person = gl(n = 3,
k = no.days,
labels = c("non-smoker",
"moderate smoker",
"heavy smoker")))Applied Machine Learning and Predictive Modelling 1 - Notes
Week 01: Linear Models
Regression
\[ y = \beta_0 + \beta_1 \cdot x_1 + \epsilon \]
\(\beta_0\) and \(\beta_1\) are the regression parameters:
- \(\beta_0\) is also the intercept
- \(\beta_1\) ist the slope
Coefficients
The coefficients are estimated from data. Estimated regression coefficients are denoted with a hat (e.g. \(\hat{\beta_1}\)). Fitted values (i.e. what the model predicts) are denoted with a hat as well (i.e. \(\hat{y}\)). Residuals are the difference between observed and predicted values.
\[ \text{res} = y - \hat{y} \]
If the errors are normally distributed, regression coefficients can be tested with t-tests
Dichotomising p-values into ”significant”/”non-significant” is very bad practice!
The grade of fit can be quantified with \(R^2\)
\[ R^2 = corr(y, \hat{y})^2 \]
Note: \(R^2\) is not used to formally compare model.
p-values
The p-value quantifies the probability of observing the value of the test statistic, or a more extreme value, under the null hypothesis. Low p-values are coherent with a rejection of the null hypothesis stating that there is no effect. Large p-values (close to 1) do not imply the we can accept the null hypothesis.
Testing
Categorical Values
Categorical values can be tested via F-tests by using the drop1() function or by comparing two models via the anova() function. Comparisons among levels of a factor (i.e. ”contrasts”) can be performed by using the glht() function.
Continuous or discrete variables
Continuous (and discrete) variables can be tested via F-tests (with drop1()) or by t-tests (with summary()). Sometimes the inferential results for continuous variables are best displayed and communicated with confidence intervals.
Interactions
If an interaction term is shown to be significant, then all terms involved in this interaction play a relevant role. An interaction involving two predictors is called ”two-fold interaction” (e.g. age * species)
Note: An interaction can involve more than two predictor.
Week 02: Modelling non-linearities
A Model is called to be linear if it is linear on its coeffitinents. By including polynomials (e.g. \(x_1 + {x_1}^2\)) we can model non-linear relationships with a linear model
The term “linear” here does not refer to the shape of the curve you draw in the coordinate system, but to the way the parameters (the coefficients \(\beta\)) appear in the equation.
Non-linear effect
Linear models can model non-linear effects!
Non-linear models are non-linear in their coefficients.
\[ y = \beta_0 + \beta_1 \cdot x^{\beta_2} + \epsilon \]
Generalise Additive Models (GAM)
GAMs come with advantages and disadvantages compared to e.g. polynomials:
Advantages
- the degree of complexity is automatically chosen
- the ”estimated degrees of freedom” give the user an indication of the complexity of a given smooth term
- smooth terms can be visualised
Disadvantages
- GAMs can run into computational issues (e.g. models that do not converge)
- the use of a quadratic term is simpler to explain than a GAM to a non-technical audience
- in order to fit and understand the results of a GAM some technical knowledge is required
Week 03: Generalised Linear Models
Linear Model Assumptions
The inference on the regression coefficients (p-values and CIs) is based on the following assumption about the errors:
\[ \varepsilon \stackrel{\text{iid}}{\sim} \mathcal{N}(0, \sigma^2) \]
This means that:
- The errors follow a normal distribution
- The errors expected value is zero
- The errors are homoscedastic (constant variance)
- The errors are independent
Count Data
Below, we list a few examples of count data:
- Items purchased by one client during a single shopping session.
- Number of children in a family.
- Faulty parts found in a car during an inspection.
Modelling count data with a linear model
If we were to use a linear model to model these data, we would assume the data to follow a normal distribution.
Note: I added a single cigarette to the non-smoker because if you don’t do so all p-values are 1
lm.smokers <- lm(no.cigarettes ~ person, data = d.smokers)
round(coef(lm.smokers), digits = 1) (Intercept) personmoderate smoker personheavy smoker
0.1 5.0 20.8 The estimates of this linear model, which assumes normality of the data, look good. Let’s dig a bit further and simulate some new observations from this linear model.
library(ggplot2)
set.seed(3)
sim.data.smokers <- simulate(lm.smokers)
ggplot(
mapping = aes(
y = sim.data.smokers$sim_1,
x = d.smokers$person)) +
geom_boxplot() +
geom_hline(yintercept = 0) +
ylab("simulated no. of cigarettes\n(assuming normality)") +
xlab("person")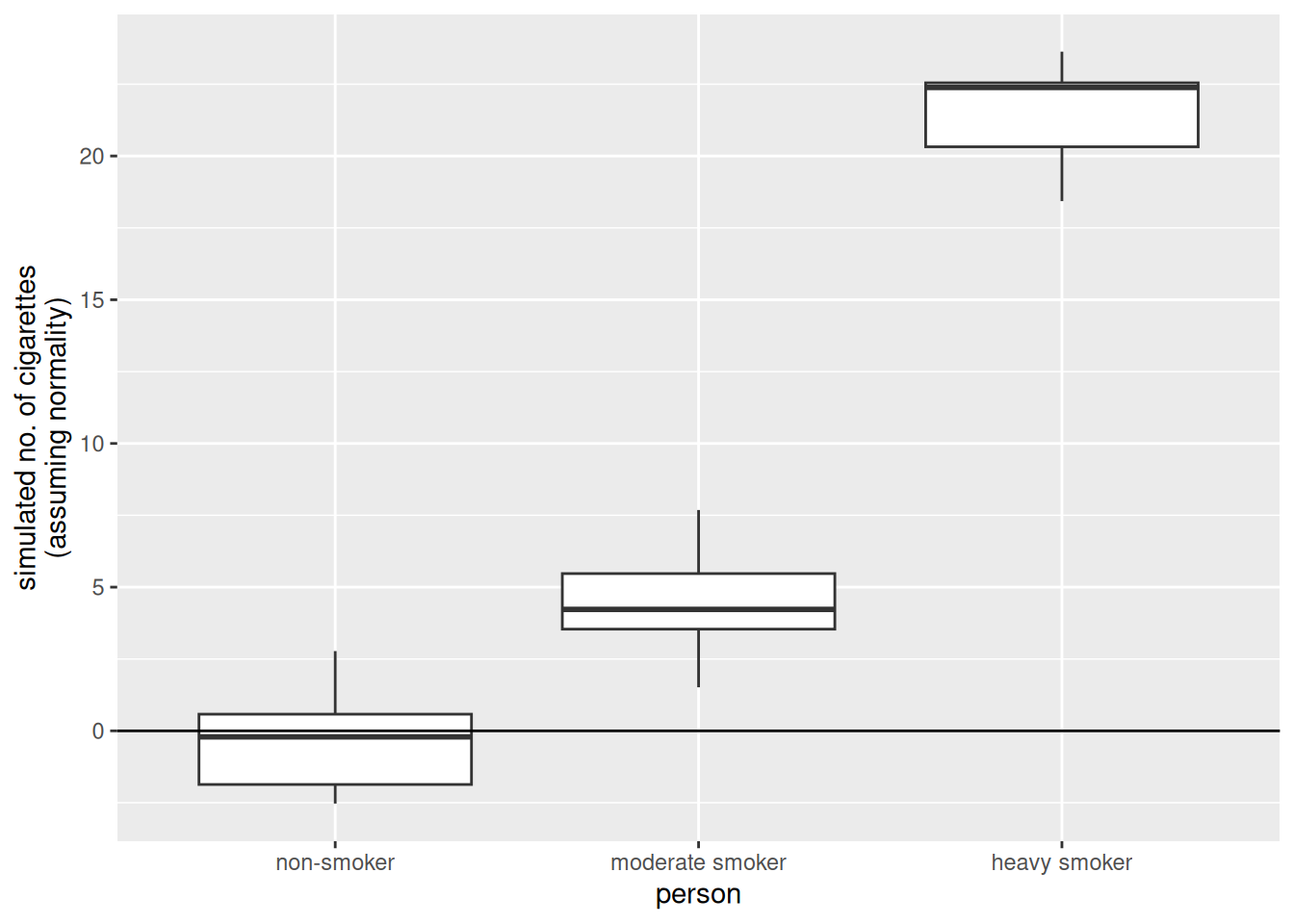
There are some striking differences between the data we simulated from the linear model and the data that made up:
- The made up data are integers, the data simulated from the linear model are numeric (note that none of the simulated values is an integer).
- The made up data are always larger than zero, the data simulated from the linear model go below zero as well (i.e., a negative number of cigarettes)
- The variability in the made up data increases with the mean value of the person (i.e., “variability non smoker” < “variability moderate smoker” < “variability heavy smoker”). On the other hand, the variability of the simulated values is the same in all three groups.
The Poisson model
Let’s see how we could modify the linear model such that these requirements that come from the very nature of count data are fulfilled.
Avoiding negative fitted values
Consider the equation we have used so far when computing the fitted values of a linear model.
\[ \hat{y} = \hat{\beta}_0 + \hat{\beta}_1 \cdot x_1 \]
To solve the problem of negative fitted values, we must make sure that the right hand-side of this equation always produces positive results. One solution is to apply a transformation to it. For example, we can use the exponential function.
\[ \hat{y} = \exp(\hat{\beta}_0 + \hat{\beta}_1 \cdot x_1) \]
Note: Ee can rewrite the above equation by taking the logarithm on both sides.
Integers and non-constant variance
we also need to make sure that the simulated values are integers and that the variance of the observations depends on the mean values. For count data, the classical solution is to assume the data to follow a Poisson distribution (rather than the normal distribution).
\[ \log(\hat{y}) = \hat{\beta}_0 + \hat{\beta}_1 \cdot x_1 | \hat{y} = \text{Poisson}(\lambda) \]
Note: This model fullfills all the three requirements we listed above.
As a matter of fact, for the Poisson distribution the variance of the observations linearly increases with the mean value.
Generalised Linear Models
Linear models are well suited to very many situations encountered in practice. However, they are not suited when the response variable represents:
- Count data: There the Poisson Model is more adequate
- Binary or binomial data: There the Binomial Model is more adequate
Both these models are an extension of the linear model These models belong to the ”class” of Generalised Linear Models GLMs generalise the linear model such that non-normal data can be analysed.
Note that in practice count and binomial data are virtually always overdispersed.
The Poisson model in R
To fit a Poisson model in R we can use the glm() function, where we specify the distribution to be used. GLMs generalise the linear model in the sense that the data can follow distributions other than normal.
glm.smokers <- glm(
no.cigarettes ~ person,
family = "poisson", ## we specify the distribution!
data = d.smokers
)Let’s simulate some data from this model.
set.seed(2)
sim.data.smokers.Poisson <- simulate(glm.smokers)
ggplot(
mapping = aes(
y = sim.data.smokers.Poisson$sim_1,
x = d.smokers$person
)
) +
geom_boxplot() +
geom_hline(yintercept = 0) +
ylab("simulated no. of cigarettes\n(assuming Poisson dist)") +
xlab("person")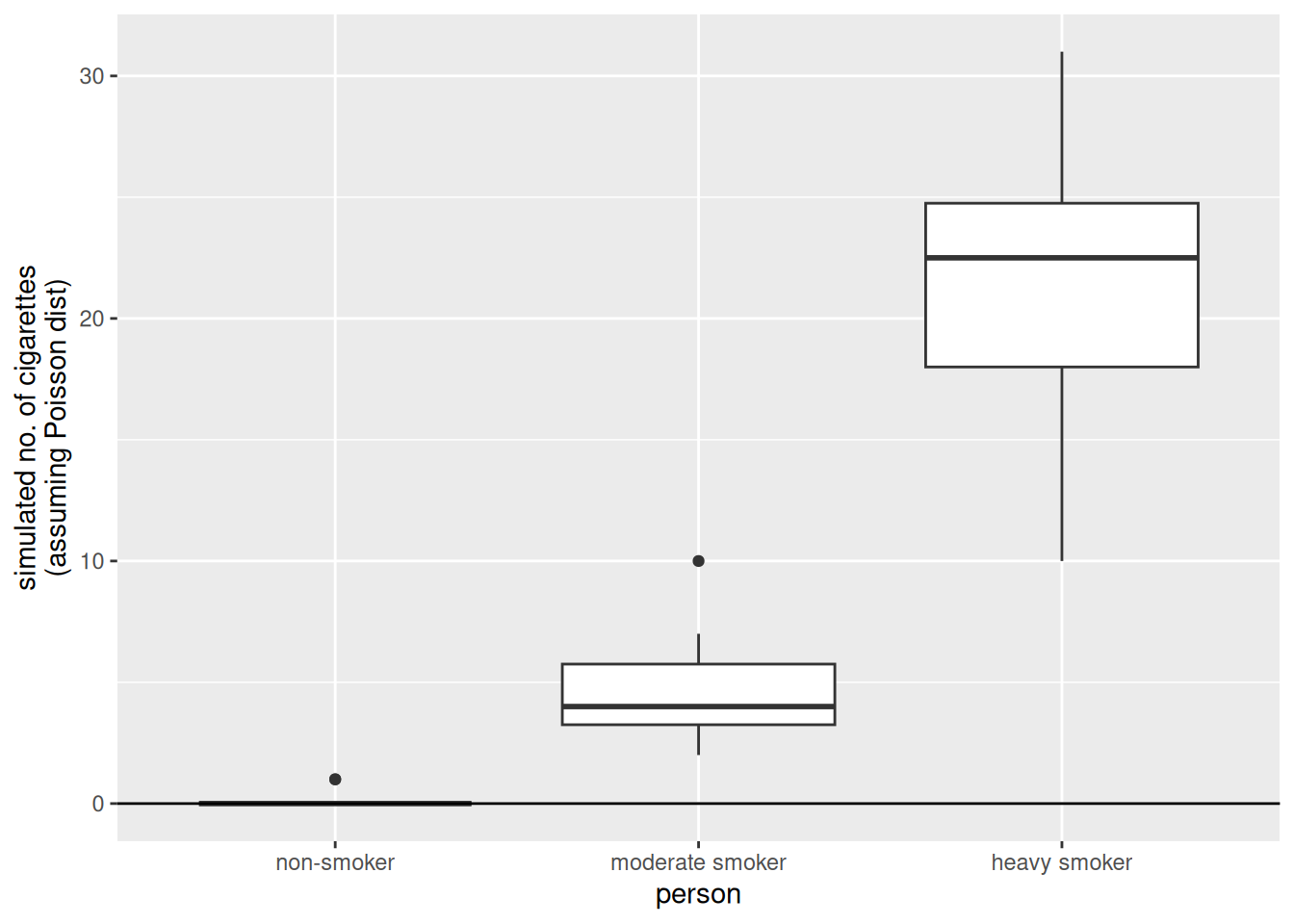
The results obtained simulating from the Poisson model are much more similar to the observations.
Binary and binomial data
Examples
Binary data (0 or 1):
- Whether a client visiting a website clicks on advertisement banner.
- Whether a cancer patient survived an “invasive” surgery.
Binomial data (number of success / number of trials) \(\in [0, 1]\):
- The proportion of students that passes an exam (e.g., hopefully all).
- The proportion of trains that gets to destination on time.
The bliss data set
library(faraway)
data(bliss)
library(dplyr)
Attaching package: 'dplyr'The following objects are masked from 'package:stats':
filter, lagThe following objects are masked from 'package:base':
intersect, setdiff, setequal, unionbliss <- bliss %>%
mutate(mortality.rate = round(dead / (dead + alive), digits = 2))
ggplot(
data = bliss,
mapping = aes(
y = mortality.rate,
x = conc
)
) +
geom_point() +
geom_smooth(method = "lm", se = FALSE) +
ylim(0, 1) +
geom_hline(yintercept = 0:1, colour = "grey")`geom_smooth()` using formula = 'y ~ x'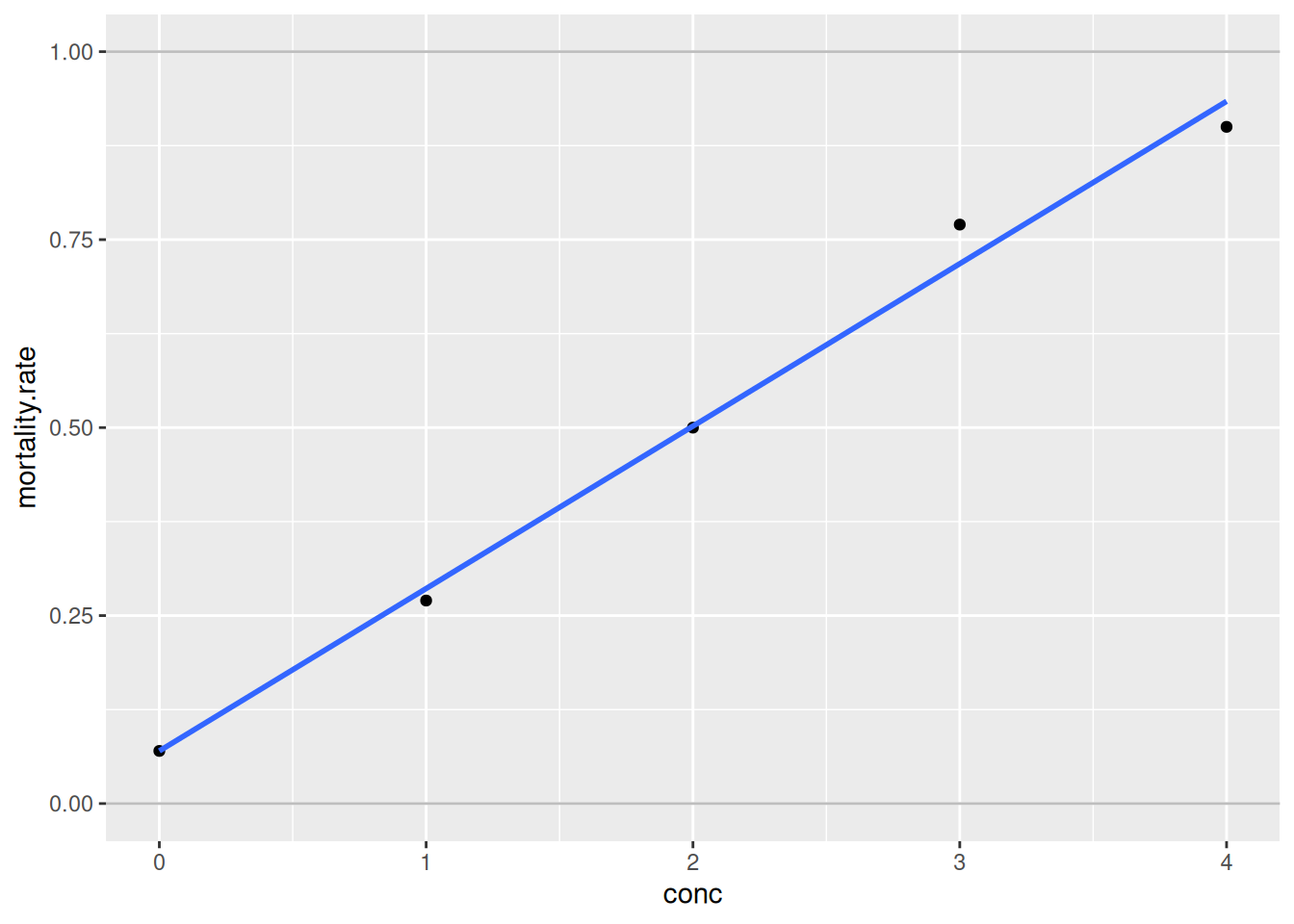
The fit seems to be fairly good. However, what problem could arise when fitting a linear model to these data?
- Predictions may lay outside the probability range [0%, 100%].
- Even fitted values may lay outside [0%, 100%].
- Actually, the response variable is not normally distributed. Its distribution is bounded between 0% and 100%. The normal distribution is unbounded and thus inconvenient here.
- The variability of the observations is not constant as it is assumed by the normal distribution. This is not evident on the graph above as there is only one observation for each concentration group. If there were many observations per group, we could see that the variability is not constant along the y-axis.
Link function for binomial data
The solution to problems 1. and 2. is to use a link function such that predictions are constrained in the [0%, 100%] range. For binary or binomial data the default choice of the link function is the “logit”.
\[ \text{logit}(y) = \log(\dfrac{y}{1-y}) | y \in [0, 1] \]
Once you applied the link function to the left-hand side of this equation, you can bring it to the right-hand side by taking its inverse on both sides.
\[ \hat{y} = \text{inverse.logit}(\hat{\beta}_0 + \hat{\beta}_1 \cdot x_1) \]
To deal with the peculiarities of count and binomial/binary data, we use a link function and assume a distribution other than the normal. This is the essence of GLMs.
The Binomial model in R
Let’s fit a Binomial model to the bliss dataset.
glm.insects <- glm(
cbind(dead, alive) ~ conc,
family = "binomial",
data = bliss
)Let’s visualise the results of this model.
new.data = data.frame(conc = seq(0, 5, length.out = 100))
new.data$pred.insects <- predict(glm.insects, newdata = new.data,
type = "response")
##
ggplot(data = bliss,
mapping = aes(y = mortality.rate,
x = conc)) +
ylim(0,1) +
geom_hline(yintercept = 0:1, col = "gray") +
##
## predictions for conc 0 --> 5
geom_point(data = new.data,
mapping = aes(
y = pred.insects,
x = conc)) +
##
## actual observations
geom_point(col = "red",
size = 3)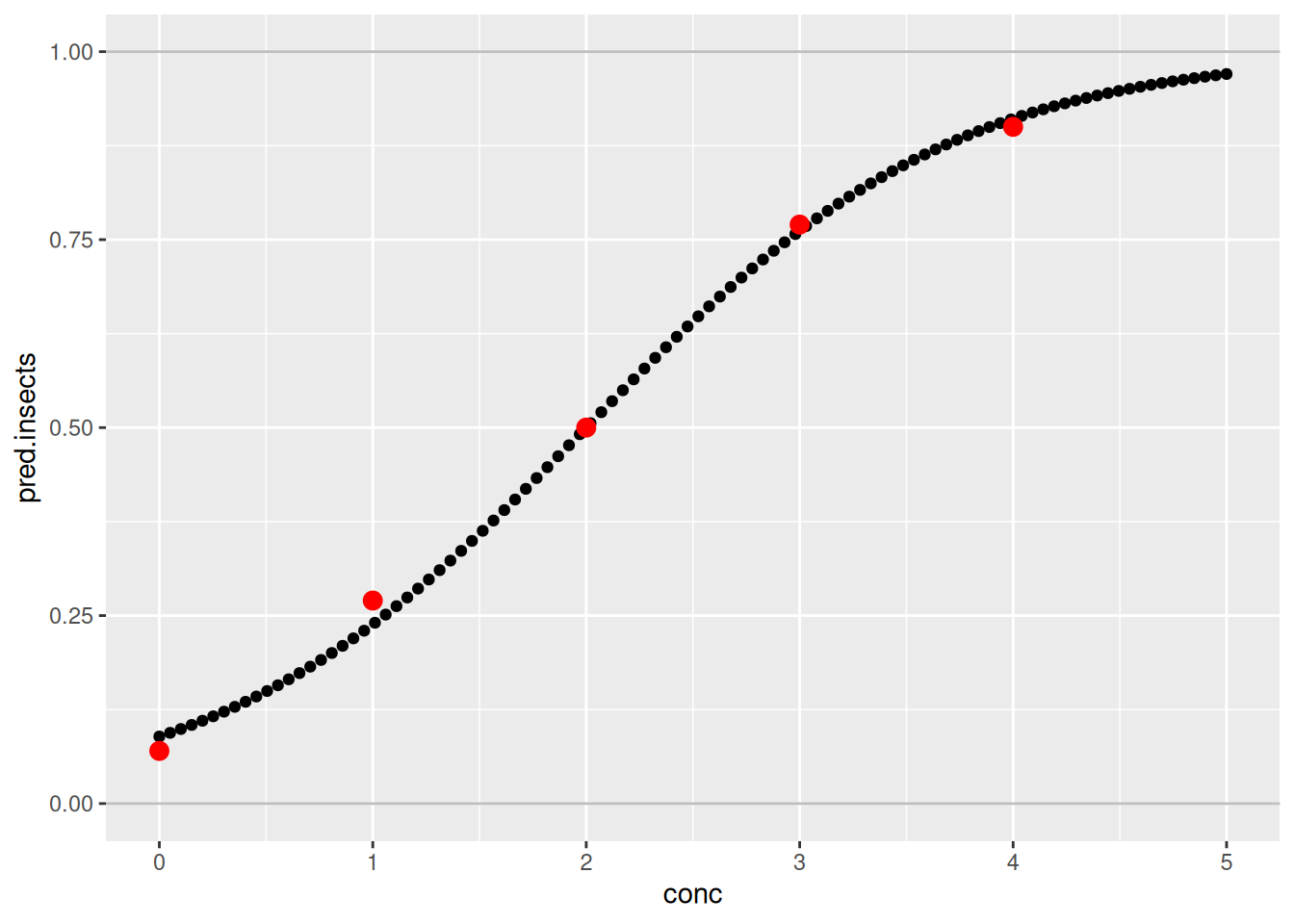
The fit of the model is quite good. Indeed, the predictions are quite close to the real observations.
Further on binary and binomial data
In many real applications there are more than two possible outcomes. Fortunately, an extension of the logistic regression to more than two classes exist. This method is called multinomial regression and is implemented in the multinom() function from the {nnet} package.
library(nnet)
multinom.iris <- multinom(
Species ~ Sepal.Length + Petal.Width,
trace = FALSE,
data = iris)Estimating the performance of a binary model
A very common way to estimate the performance of a binary model is too look at predicted values and compare them with the actual observations.
d.stab <- read.table(
"/home/nils/dev/mscids-notes/fs26/mpm1/data/stability.dat",
stringsAsFactors = TRUE,
header = TRUE
)
str(d.stab)'data.frame': 27 obs. of 2 variables:
$ perform : num 0 0 0 1 1 0 0 1 1 1 ...
$ stability: num 474 432 453 481 619 584 399 582 638 624 ...ggplot(
data = d.stab,
mapping = aes(
y = perform,
x = stability
)
) +
geom_point()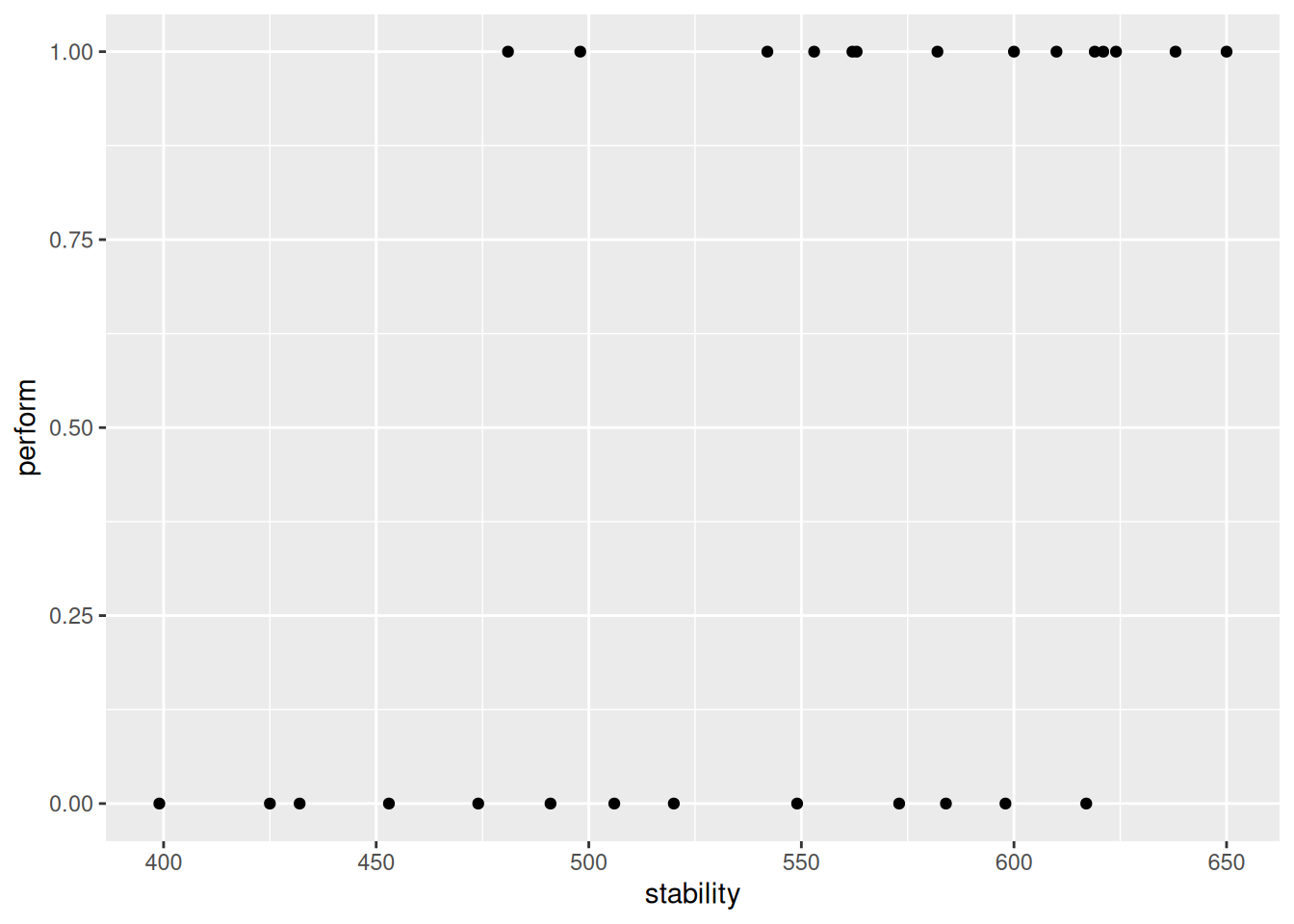
There are only two possible outcomes here: success or failure. Let’s fit a logistic regression model and add the fit to the this graph.
ggplot(
data = d.stab,
mapping = aes(
y = perform,
x = stability
)
) +
geom_point() +
geom_smooth(
method = "glm",
se = FALSE,
method.args = list(family = "binomial")
) `geom_smooth()` using formula = 'y ~ x'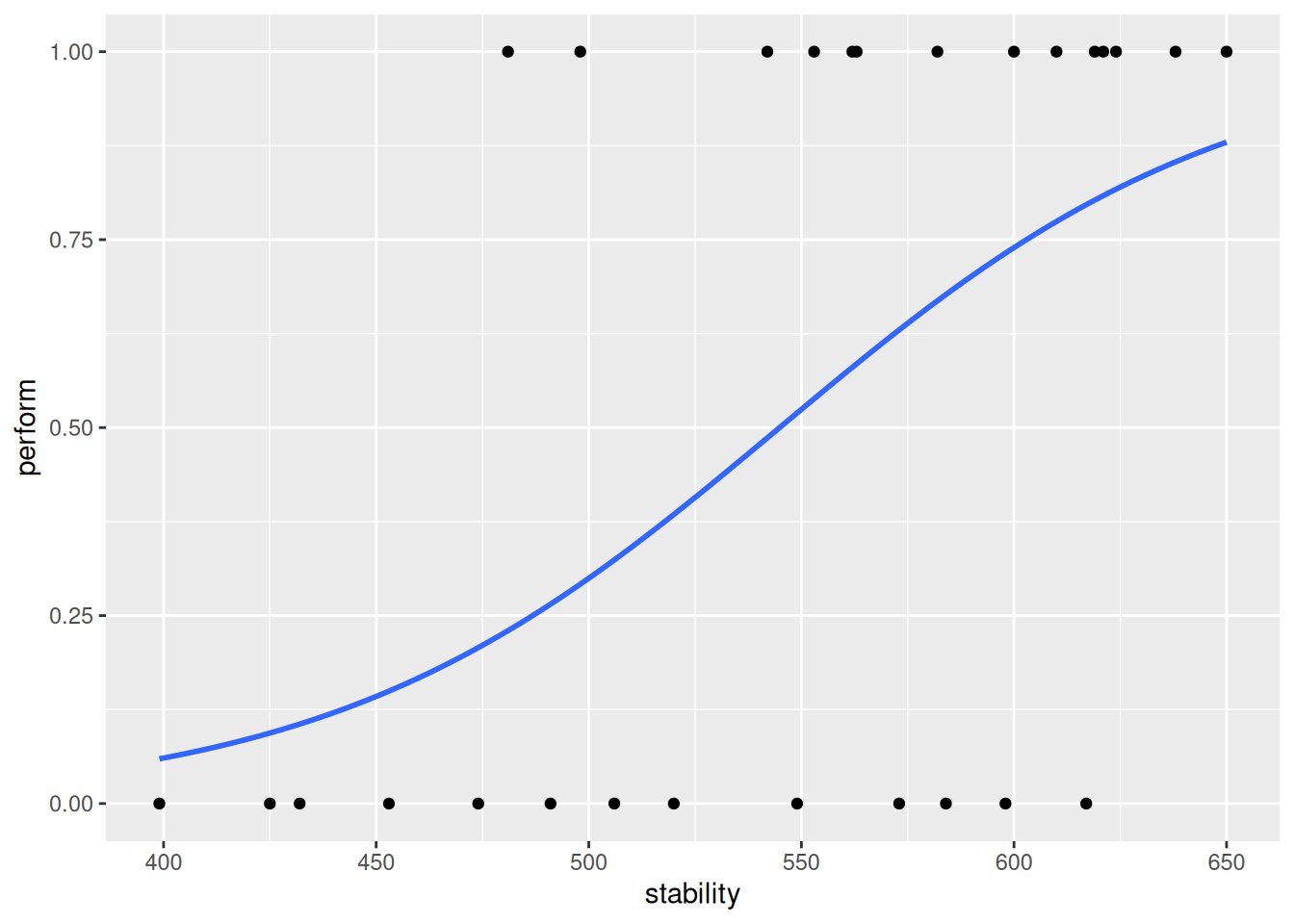
As expected, stability has a positive effect on the response variable. Let’s have a look at the fitted values for this model.
glm.stab <- glm(
perform ~ stability,
data = d.stab,
family = "binomial")
fitted(glm.stab) %>%
round(digits = 2) 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16
0.21 0.11 0.15 0.23 0.80 0.68 0.06 0.67 0.85 0.82 0.49 0.88 0.54 0.09 0.58 0.52
17 18 19 20 21 22 23 24 25 26 27
0.29 0.38 0.77 0.73 0.27 0.80 0.81 0.63 0.58 0.32 0.74 Note: They are probabilities, thus, bounded within 0 and 1.
To compare the fitted values with the observed values, which are indeed binary (0 or 1), we can also discretise/dichotomise the fitted values into 0 and 1. In order to do that, we use a cutoff of 0.5. Finally, let’s compare the observed and fitted values.
fitted.stab.disc <- ifelse(
fitted(glm.stab) < 0.5,
yes = 0,
no = 1
)
d.obs.fit.stab <- data.frame(
obs = d.stab$perform,
fitted = fitted.stab.disc)
table(
obs = d.obs.fit.stab$obs,
fit = d.obs.fit.stab$fitted) fit
obs 0 1
0 8 5
1 3 11The diagonal entries of this matrix represent correctly labelled observations
Week 04: Support Vector Machines
What is an SVM?
Support-vector machines (SVMs, also support-vector networks) are supervised learning models with associated learning algorithms that analyze data used for classification and regression analysis. An SVM model is a representation of the examples as points in space, mapped so that the examples of the separate categories are divided by a clear gap that is as wide as possible.
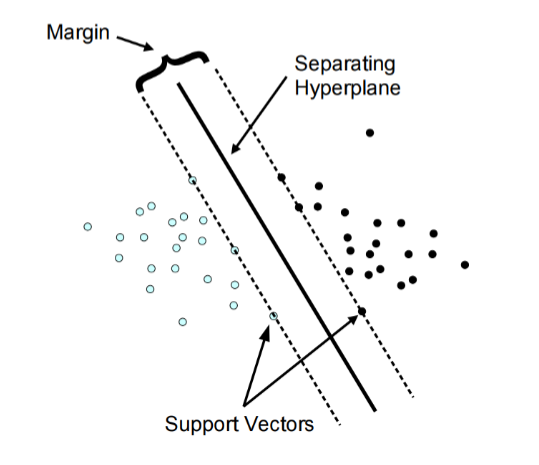
set.seed(123)
library(tidyverse)── Attaching core tidyverse packages ──────────────────────── tidyverse 2.0.0 ──
✔ forcats 1.0.1 ✔ stringr 1.6.0
✔ lubridate 1.9.4 ✔ tibble 3.3.1
✔ purrr 1.2.1 ✔ tidyr 1.3.2
✔ readr 2.1.6
── Conflicts ────────────────────────────────────────── tidyverse_conflicts() ──
✖ purrr::%||%() masks base::%||%()
✖ dplyr::filter() masks stats::filter()
✖ dplyr::lag() masks stats::lag()
ℹ Use the conflicted package (<http://conflicted.r-lib.org/>) to force all conflicts to become errorslibrary(e1071)
Attaching package: 'e1071'
The following object is masked from 'package:ggplot2':
elementlibrary(caret)Loading required package: lattice
Attaching package: 'lattice'
The following object is masked from 'package:faraway':
melanoma
Attaching package: 'caret'
The following object is masked from 'package:purrr':
lift# Load data
data(iris)
# Prepare the Data for Training
indices <- createDataPartition(iris$Species, p=.85, list = F)
train <- iris %>%
slice(indices)Warning: Slicing with a 1-column matrix was deprecated in dplyr 1.1.0.test_in <- iris %>%
slice(-indices) %>%
select(-Species)
test_truth <- iris %>%
slice(-indices) %>%
pull(Species)
# Train the Support Vector Machine
iris_svm <- svm(Species ~ ., train, kernel = "linear", scale = TRUE, cost = 10)
plot(iris_svm, train, Petal.Length ~ Petal.Width)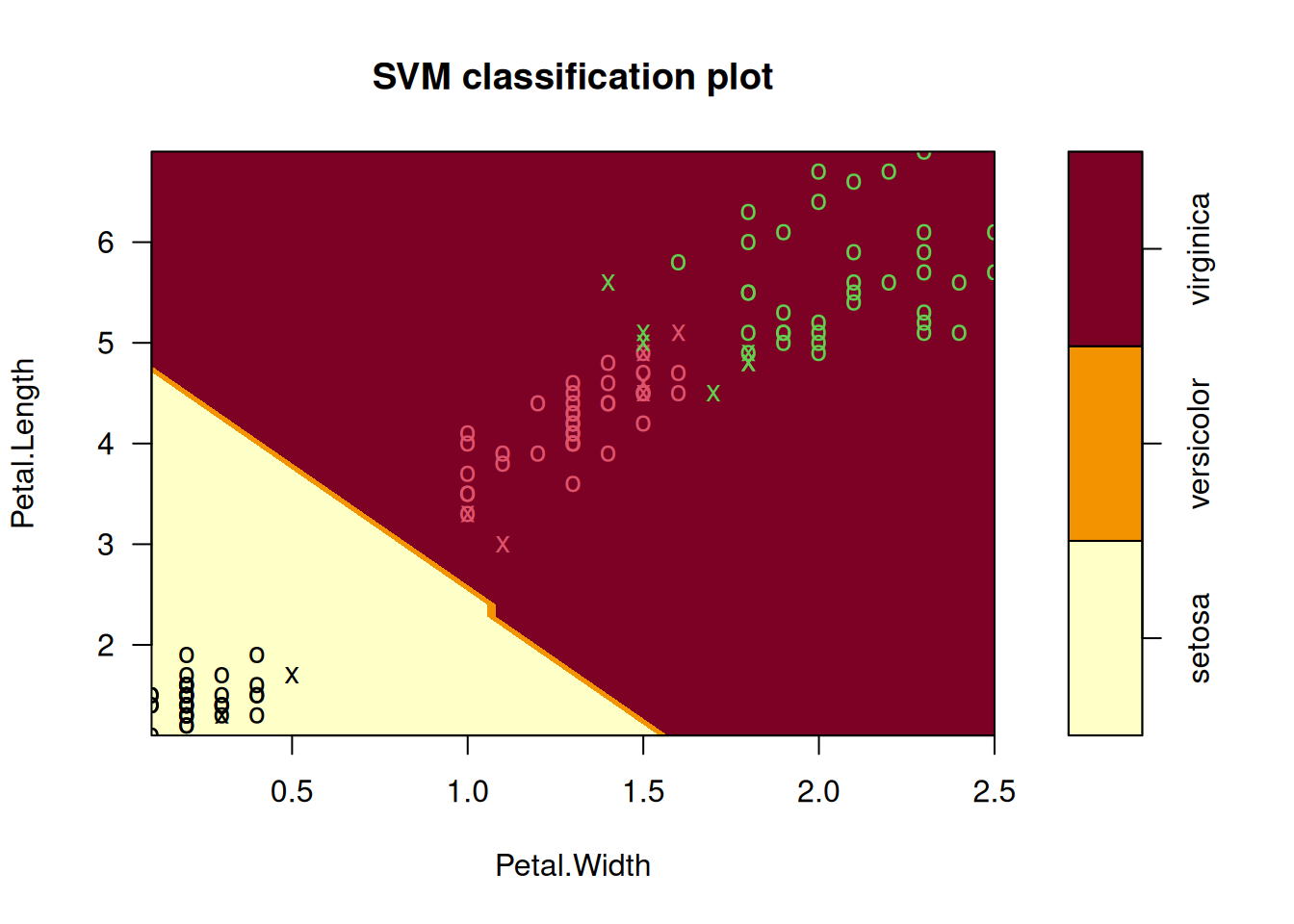
SVM Kernel Functions
In many real-world scenarios, data is non-linearly separable. A standard linear SVM would fail here. Instead of manually performing complex mathematical transformations to move data into a higher dimension (which is computationally expensive), we use the Kernel Trick. A kernel function calculates the “similarity” or “inner product” between two points in a high-dimensional space without actually transforming the data points themselves.
Soft Margin Formulation
If you try to force a Hard Margin SVM to separate every single point perfectly, you often end up with overfitting or a model that simply fails to find a solution because the data overlaps.
The Soft Margin Formulation was introduced to solve this by allowing the SVM to make a few “mistakes” for the sake of a better, more general boundary.
The Role of the Hyperparameter \(C\)
The variable \(C\) is the most important dial you can turn. it controls the trade-off between “smoothness” and “accuracy.”
Analyse Model Performance
test_pred <- predict(iris_svm, test_in)
table(test_pred)test_pred
setosa versicolor virginica
7 6 8 conf_matrix <- confusionMatrix(test_pred, test_truth)
conf_matrixConfusion Matrix and Statistics
Reference
Prediction setosa versicolor virginica
setosa 7 0 0
versicolor 0 6 0
virginica 0 1 7
Overall Statistics
Accuracy : 0.9524
95% CI : (0.7618, 0.9988)
No Information Rate : 0.3333
P-Value [Acc > NIR] : 4.111e-09
Kappa : 0.9286
Mcnemar's Test P-Value : NA
Statistics by Class:
Class: setosa Class: versicolor Class: virginica
Sensitivity 1.0000 0.8571 1.0000
Specificity 1.0000 1.0000 0.9286
Pos Pred Value 1.0000 1.0000 0.8750
Neg Pred Value 1.0000 0.9333 1.0000
Prevalence 0.3333 0.3333 0.3333
Detection Rate 0.3333 0.2857 0.3333
Detection Prevalence 0.3333 0.2857 0.3810
Balanced Accuracy 1.0000 0.9286 0.9643Pros & Cons of SVM
Pros
- It works really well with a clear margin of separation
- It is effective in high dimensional spaces
- It is effective in cases where the number of dimensions is greater than the number of samples
- It uses a subset of training points in the decision function (support vectors), so it is memory efficient
Cons
- It is not always straightforward to scale the different variables “equally”
- It is very prone to overfitting (different Kernel functions, hyperparameters etc.)
- It doesn’t perform well when we have large data set because the required training time is higher
- It also doesn’t perform very well, when the data set has more noise i.e. target classes are overlapping
- It doesn’t directly provide probability estimates
Support Vector Regression
SVR gives us the flexibility to define how much error is acceptable in our model and will find an appropriate line (or hyperplane in higher dimensions) to fit the data. In contrast to OLS, the objective function of SVR is to minimize the coefficients not the squared error.
Summary
- Support Vector Machine (SVM) is a non-probabilistic binary linear classifier
- It can handle non-linear boundaries with the Kernel function/trick
- It can be extended to more than two classes as well
- It is also possible to do regression using this technique (Support Vector Regression)
- Particularly useful in cases where we want to ignore errors up to a certain tolerance
- Be careful with scaling your data and overfitting!
Week 05: Model Validation
The validation should simulate the setting in which the model is used. The choice of the most suited method depends on the situation. ften several methods are used simultaneously (to compare results).
Data Set Approaches
Validation Set Approach
A set of \(n\) observations is randomly split into training and validation sets. The statistical learning method is then fitted to the training set and its performance evaluated on the validation set.
set.seed(1)
df <- data.frame(
x = 1:100,
y = (1:100) * 2 + rnorm(100)
)
n <- nrow(df)
train_size <- floor(0.7 * n)
train_indices <- sample(seq_len(n), size = train_size)
train_set <- df[train_indices, ]
test_set <- df[-train_indices, ]
train_set x y
77 77 153.556708
90 90 180.267099
98 98 195.426735
66 66 132.188792
19 19 38.821221
17 17 33.983810
34 34 67.946195
75 75 148.746367
31 31 63.358680
35 35 68.622940
46 46 91.292505
96 96 192.558486
93 93 187.160403
16 16 31.955066
40 40 80.763176
9 9 18.575781
50 50 100.881108
24 24 46.010648
10 10 19.694612
79 79 158.074341
32 32 63.897212
39 39 79.100025
37 37 73.605710
12 12 24.389843
14 14 25.785300
87 87 175.063100
15 15 31.124931
2 2 4.183643
65 65 129.256727
67 67 132.195041
95 95 191.586833
5 5 10.329508
41 41 81.835476
92 92 185.207868
36 36 71.585005
49 49 97.887654
68 68 137.465555
29 29 57.521850
21 21 42.918977
1 1 1.373546
53 53 106.341120
20 20 40.593901
42 42 83.746638
99 99 196.775387
58 58 114.955865
83 83 167.178087
60 60 119.864945
54 54 106.870637
22 22 44.782136
38 38 75.940687
84 84 166.476433
88 88 175.695816
30 30 60.417942
63 63 126.689739
73 73 146.610726
8 8 16.738325
76 76 152.291446
97 97 192.723408
26 26 51.943871
86 86 172.332950
78 78 156.001105
45 45 89.311244
7 7 14.487429
43 43 86.696963
64 64 128.028002
85 85 170.593946
27 27 53.844204
74 74 147.065902
25 25 50.619826
6 6 11.1795325-fold CV
A set of \(n\) observations is randomly divided into five non-overlapping groups. Each of these fifths acts as a validation set and the remainder as a training set. The test error is estimated by averaging the five resulting mean squared error (MSE) estimates.
set.seed(1)
df <- data.frame(x = 1:100, y = rnorm(100))
df <- df[sample(nrow(df)), ]
folds <- cut(seq(1, nrow(df)), breaks = 5, labels = FALSE)
test_indices <- which(folds == 1)
test_data <- df[test_indices, ]
train_data <- df[-test_indices, ]
train_data x y
32 32 -0.102787727
39 39 1.100025372
37 37 -0.394289954
12 12 0.389843236
14 14 -2.214699887
87 87 1.063099837
15 15 1.124930918
2 2 0.183643324
65 65 -0.743273209
67 67 -1.804958629
95 95 1.586833455
5 5 0.329507772
41 41 -0.164523596
92 92 1.207867806
36 36 -0.414994563
49 49 -0.112346212
68 68 1.465554862
29 29 -0.478150055
21 21 0.918977372
1 1 -0.626453811
53 53 0.341119691
20 20 0.593901321
42 42 -0.253361680
99 99 -1.224612615
58 58 -1.044134626
83 83 1.178086997
60 60 -0.135054604
54 54 -1.129363096
22 22 0.782136301
38 38 -0.059313397
84 84 -1.523566800
88 88 -0.304183924
30 30 0.417941560
63 63 0.689739362
73 73 0.610726353
8 8 0.738324705
76 76 0.291446236
97 97 -1.276592208
26 26 -0.056128740
86 86 0.332950371
78 78 0.001105352
45 45 -0.688755695
7 7 0.487429052
43 43 0.696963375
64 64 0.028002159
85 85 0.593946188
27 27 -0.155795507
74 74 -0.934097632
25 25 0.619825748
6 6 -0.820468384
48 48 0.768532925
71 71 0.475509529
4 4 1.595280802
3 3 -0.835628612
94 94 0.700213650
13 13 -0.621240581
91 91 -0.542520031
51 51 0.398105880
44 44 0.556663199
69 69 0.153253338
23 23 0.074564983
82 82 -0.135178615
81 81 -0.568668733
61 61 2.401617761
100 100 -0.473400636
52 52 -0.612026393
72 72 -0.709946431
80 80 -0.589520946
33 33 0.387671612
89 89 0.370018810
57 57 -0.367221476
70 70 2.172611670
55 55 1.433023702
28 28 -1.470752384
59 59 0.569719627
62 62 -0.039240003
18 18 0.943836211
11 11 1.511781168
47 47 0.364581962
56 56 1.980399899Leave one out CV (LOOCV)
A set of \(n\) data points is repeatedly split into a training set containing all but one observation and a validation set containing only that observation. The test error is estimated by averaging the resulting MSE’s.
set.seed(1)
df <- data.frame(x = 1:100, y = rnorm(100))
n <- nrow(df)
predictions <- numeric(n)
# The LOOCV Loop
for (i in 1:n) {
train_data <- df[-i, ]
test_data <- df[i, ]
model <- lm(y ~ x, data = train_data)
predictions[i] <- predict(model, test_data)
}
mse_loocv <- mean((df$y - predictions)^2)
mse_loocv[1] 0.8317288Week 06: Artificial Neural Networks
Artificial neural networks (ANNs), usually simply called neural networks (NNs), are computing systems vaguely inspired by the biological neural networks that constitute animal brains. An ANN is based on a collection of connected units or nodes called artificial neurons, which loosely model the neurons in a biological brain. Each connection, like the synapses in a biological brain, can transmit a signal to other neurons. An artificial neuron that receives a signal then processes it and can signal neurons connected to it.
Motivation & History
Model a program that works similar to the Human Brain. Build of different neurons (up to \(10^9\)). Type and strength of connection influence the flow of the signals.
The Perceptron
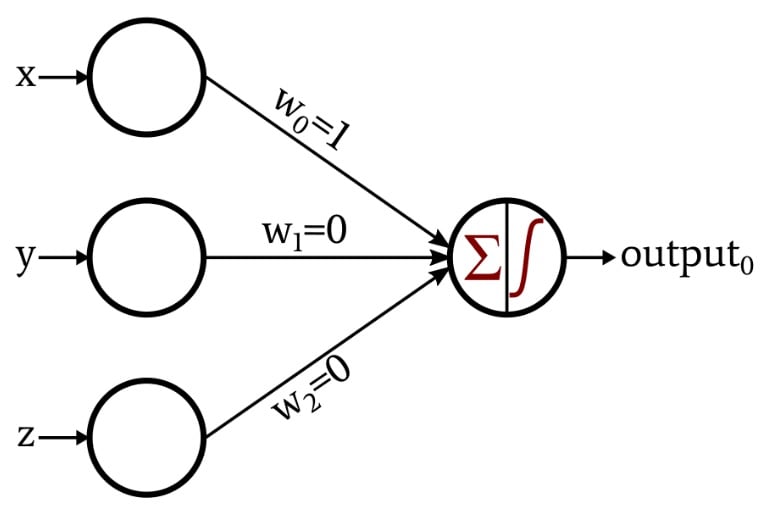
A Perceptron is an algorithm for supervised learning of binary classifiers. It mimics a single biological neuron by receiving multiple signals, processing them, and deciding whether or not to “fire” an output.
- Input Values (\(x_n\)): These are the features of your data (e.g., pixels in an image or dimensions of a fruit).
- Weights (\(w_n\)): Each input is assigned a weight that represents its importance. The network “learns” by adjusting these weights.
- Summation Function: The Perceptron calculates the weighted sum of the inputs and adds a bias (\(b\)). The bias allows the model to shift the activation function up or down.
Activation Functions
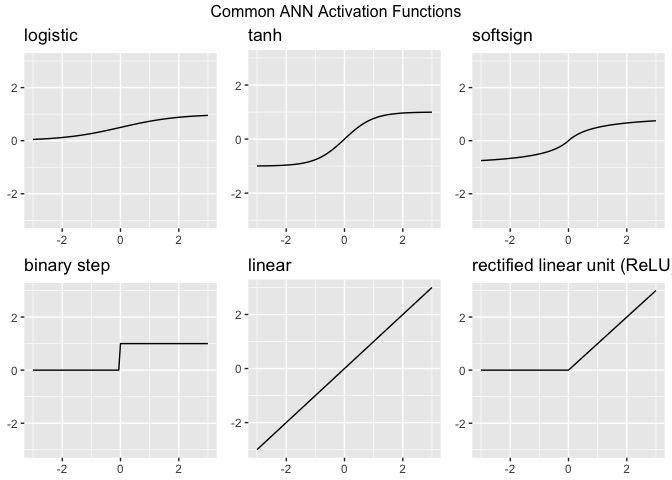
Activation functions are the “gatekeepers” of a neural network. While the Perceptron uses a simple step function, modern Artificial Neural Networks (ANNs) use more complex functions to help the model learn intricate patterns. Without an activation function, a neural network is just a giant linear regression model—no matter how many layers you add, it could only represent linear relationships. Activation functions introduce non-linearity, allowing the network to learn “curvy” and complex boundaries.
Layers
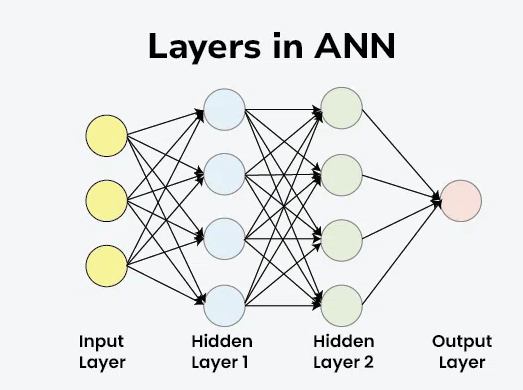
Layers are the structural building blocks that organize how data is processed. Instead of just one neuron (like the Perceptron), we stack them to create “Deep” networks. Think of layers as a filtration system: each layer extracts more complex features from the raw data until the final layer can make a confident prediction.
- Input Layer: Receive data
- Hidden Layer: Feature extraction & pattern recognition
- Output Layer: Final prediction / Classification
As you add more hidden layers, it becomes harder for humans to understand exactly why a network made a specific decision. This is why deep learning is often referred to as a “black box.”
Gradient Descent
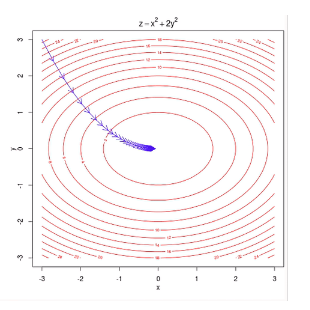
Gradient Descent is an iterative optimization algorithm used to find the minimum of a function. In the context of ANNs, that function is the Loss Function (the measure of how “wrong” the model’s predictions are).
Learning Rate
The size of the “steps” you take down the mountain is controlled by the Learning Rate. This is a crucial “hyperparameter” you set before training.
- Too High: You might overstep the valley and bounce back and forth, never reaching the bottom.
- Too Low: It will take forever to reach the bottom, and you might get stuck in a “Local Minimum” (a small dip that isn’t the true lowest point).
Deep Learning
Deep learning is essentially the “heavy lifting” department of Artificial Intelligence. While traditional AI relies on human-coded rules, deep learning uses Artificial Neural Networks to mimic how the human brain processes information.
Neural networks have been around since the 1950s, but they only took off recently because of two main things: Big Data (the internet provided the “textbooks” for these models) and GPU Computing (video game hardware turned out to be perfect for the complex math required).
Note: Deep Learning is just Classical Network with many hidden layers (deep).
Generative Adversial Networks (GANs)
Generative Adversarial Networks, or GANs, are a fascinating architecture in deep learning where two neural networks “fight” each other to create incredibly realistic data, such as images, music, or speech. A GAN consists of two distinct models that are trained simultaneously:
- The Generator (The Forger): Its job is to create fake data that looks real. It starts by producing random noise but learns over time which features make an image look “authentic.”
- The Discriminator (The Detective): Its job is to look at a piece of data and decide if it is “Real” (from the actual training set) or “Fake” (created by the Generator).
Week 07 - General Topics
Parametric vs. non-parametric models
Parametric Models
The word “parametric” refers to a fixed set of parameters. Before looking at any data, you decide on a specific mathematical “shape” (like a straight line). No matter how much data you throw at it, the number of parameters remains constant (e.g., just a slope and an intercept).
Non-parametric Models
The name is a bit of a misnomer—it doesn’t mean “zero parameters.” Instead, it means the number of parameters is not predetermined. The model structure grows and changes based on the data. The “parameters” are often the data points themselves or an ever-expanding set of branches and nodes.
Comparison
| Feature | Parametric Models (e.g., LM, GLM) | Non-parametric Models (e.g., Random Forest, NN) |
|---|---|---|
| Model Structure | Fixed / Predefined (e.g., a linear equation). | Flexible / Learns the shape from the data. |
| Parameter Count | Fixed (does not change with more data). | Grows (more data = more complex structure). |
| Primary Goal | Inference: Understanding the “why” and testing hypotheses. | Prediction: Achieving the highest accuracy possible. |
| Data Requirements | Works well with small datasets. | Usually requires large amounts of data. |
| Complexity | Simple, easy to interpret (“Glass Box”). | Complex, harder to explain (“Black Box”). |
| Assumptions | Strong assumptions (e.g., normality, linearity). | Minimal assumptions about the data distribution. |